New insights into the metabolic network of Methylobacterium extorquens AM1
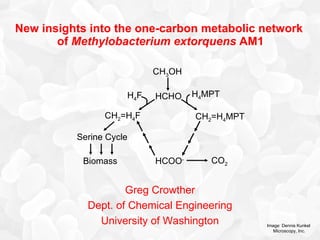
- 1. New insights into the one-carbon metabolic network of Methylobacterium extorquens AM1 Greg Crowther Dept. of Chemical Engineering University of Washington Image: Dennis Kunkel Microscopy, Inc. HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass
- 2. The “Central Dogma” of Biology DNA RNA Proteins substrates products microarrays proteomics, enzyme activity assays flux rates genomics metabolomics
- 3. My primary interest: metabolic fluxes DNA RNA Proteins substrates products flux rates
- 4. Clover leaf print showing Methylobacterium strains. Photo by Amy Springer. Methylotrophic metabolism • “ methyl” = -CH 3 , “troph” = growth • methylotrophy = growth on one-carbon compounds such as methanol (CH 3 OH ) - some methylotrophs can also grow on multicarbon compounds such as succinate (C 4 H 4 O 4 2- ) Dissimilation C 3 Assimilation C = O H H C - O H H H H CO 2 Biomass ATP NAD(P)H
- 5. • Carbon cycling Methane, a greenhouse gas, is consumed by some methylotrophs. • Bioremediation Methylotrophs detoxify many nasty compounds (e.g., chloride-containing organics). • Biocatalysis Genetic engineering enables synthesis of useful chemicals (e.g., plastics) from CH 3 OH. Images: Marina Kalyuzhnaya; appa.asso.fr; ouraycolorado.com Why is methylotrophy important? CO 2 CH 4
- 7. M ethylotrophs are somewhat widespread in domain Bacteria… Figure: Hugenholtz et al., J Bacteriol 180 : 4765, 1998 … but M. extorquens AM1 is the best-studied species • 100 + genes for one-carbon metabolism are characterized • genome is sequenced • we think we know all of the major metabolic pathways M. extorquens AM1: a model methylotroph
- 8. • pink, rod-shaped -proteobacterium • natural habitat: surface of leaves demethylation of pectin produces methanol, which is released through stomata Meet Methylobacterium extorquens AM1 Clover leaf print showing Methylobacterium strains. Photo by Amy Springer.
- 9. Metabolic fluxes in M. extorquens AM1 HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass
- 10. Dissimilation (CO 2 production) HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass
- 11. Direct assimilation pathway (Biomass production) HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass
- 12. Long assimilation pathway (Biomass production) HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass
- 13. Metabolic fluxes in M. extorquens AM1: our view as of 2005 1. The direct assimilation pathway dominates in cells growing on CH 3 OH. 2. HCHO is the key branch point (Biomass vs. CO 2 ). 3. Formate dehydrogenase (FDH) may not be important? HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass
- 14. Metabolic fluxes in M. extorquens AM1: our view as of 2005 1. The direct assimilation pathway dominates in cells growing on CH 3 OH. 2. HCHO is the key branch point (Biomass vs. CO 2 ). 3. Formate dehydrogenase (FDH) may not be important? HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass
- 15. Metabolic fluxes in M. extorquens AM1: our view as of 2005 1. The direct assimilation pathway dominates in cells growing on CH 3 OH. 2. HCHO is the key branch point (Biomass vs. CO 2 ). 3. Formate dehydrogenase (FDH) may not be important? HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass FDH
- 16. The rest of this talk: new insights, 2006-2007 1. The direct assimilation pathway: not important! HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass FDH 2. Formate dehydrogenase: very important after all!
- 17. The direct assimilation pathway Evidence for this pathway: deuterium assay (Marx et al., PLoS 3 :e16, 2005) GC-MS extract, derivatize +2 Serine ( D 2 ) +1 Serine ( D ) D COO - C D 2 =H 4 F CO 2 C D 2 =H 4 MPT H 4 MPT C D 3 O D D C D O H 4 F C D H=H 4 F 90% + of serine is +2 in CH 3 OH-grown cells, so the direct pathway appears dominant.
- 18. Potential problem with the deuterium assay If NADPH pool gets contaminated with deuterium, flux through the long pathway will be “counted” as flux through the direct pathway. GC-MS extract, derivatize +2 Serine ( D 2 ) +2 Serine ( D 2 ) D COO - C D 2 =H 4 F CO 2 C D 2 =H 4 MPT H 4 MPT C D 3 O D D C D O H 4 F C D 2 =H 4 F NADP D
- 19. If NADPH pool gets contaminated with deuterium, the +2/+1 ratio should increase as incubation time increases (and more deuterium enters the pool). Conclusion : contamination does occur and might be a problem. Testing for NADPH contamination HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass
- 20. Other reasons to question the direct pathway 1. If we knock out an enzyme in the long pathway, the cells can’t grow on CH 3 OH. Why can’t they just use the direct pathway? HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass X
- 21. Other reasons to question the direct pathway 2. We can’t find an enzyme that catalyzes HCHO + H 4 F. • Vorholt et al. ( J Bacteriol 2000): cell extracts don’t enhance reaction rate • My data ( J Bacteriol 2005): fae2 and fae3 , the genes most likely to encode the enzyme (if it exists), can be knocked out without slowing growth on CH 3 OH HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass
- 22. Other reasons to question the direct pathway 3. The nonenzymatic rate constant for HCHO + H 4 F is much lower than the deuterium and 14 C assay data would suggest. Measure 14 C-CO 2 Add 14 C-CH 3 OH Measure 14 C-Biomass HCOO - CH 2 =H 4 F Serine Cycle CH 2 =H 4 MPT H 4 MPT HCHO H 4 F
- 23. Estimates of direct pathway flux Direct pathway flux estimated from 14 C-biomass and deuterium assays = (total flux) * (direct flux / total flux) = 0.3 mM/s 14 C assay deuterium assay Is the rate constant for HCHO + H 4 F high enough to achieve this flux? HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass
- 24. rate = V 6 *[H 4 F]*[HCHO] • V 6 < 0.08 mM -1 s -1 • [H 4 F] < 0.15 mM (Vorholt et al. 1998) • [HCHO] < 1 mM rate = (0.08 mM -1 s -1 )*(0.1 mM)*(0.5 mM) = 0.004 mM/s << 0.3 mM/s Conclusion : the deuterium assay may greatly overestimate the biomass flux coming from the direct pathway. Estimates of direct pathway flux HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass
- 25. Testing the direct pathway by studying a long-pathway mutant Conclusion : Flux through the direct pathway is insignificant. If the direct pathway is “real”: • deuterium assay should detect +2 serine • 14 C assay should detect biomass flux … NO! … NO! HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass X
- 26. Can the long pathway handle the entire CH 3 OH flux? Enzyme activities from literature (converted to mM/s): 1. MDH 1.5 2. Fae 5 3. MtdA/MtdB 10-28 4. Mch 11 5. Fhc 0.2-1.1 Preliminary conclusion : Enzyme activities appear sufficient for handling all CH 3 OH. This is a question for kinetic modeling (in progress). Maximum total CH 3 OH intake = 1.4 mM/s 1 2 3 4 5 HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass
- 27. Conclusion on the direct pathway Flux through the direct assimilation pathway is insignificant. Key supporting evidence: • mutations in long pathway prevent growth on CH 3 OH • no detectable enzyme activity • rate constant for HCHO + H 4 F is very small • no +2 serine in long-pathway mutant (deuterium assay) • no biomass flux in long-pathway mutant ( 14 C assay) HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass
- 28. Formate dehydrogenase (FDH) Background : Chistoserdova et al. ( J Bacteriol 186 : 22, 2004) studied three FDH genes, each of which was shown to be functional in vivo. HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass FDH 1-3
- 29. Formate dehydrogenase (FDH) Weird findings for the triple mutant: • It still grows on CH 3 OH! • CO 2 production seems unimpaired! Possible interpretations: • There is at least one more FDH. • CO 2 is produced “downstream” of the one-carbon network. HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass FDH 1-3 X
- 30. A fourth formate dehydrogenase! A gene with sequence homology to other FDHs is upregulated in the triple mutant (E. Skovran’s microarray analysis). This gene was knocked out along with the other three. The quadruple mutant cannot grow on CH 3 OH (L. Chistoserdova), suggesting that there are 4 (and only 4) FDHs. What do the flux data show? HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass
- 31. My 14 C-CO 2 flux data CO 2 production is similar in the wild-type and triple mutant, but is virtually eliminated in the quadruple mutant. -> Consistent with the hypothesis that there are 4 and only 4 FDHs. -> Contradicts the hypothesis that there is significant CO 2 production “downstream” of the one-carbon network. HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass
- 32. HCOO - consumption data (J. Vorholt) Capacity to consume HCOO - (as measured by 13 C NMR spectroscopy), in nmol/(mg*min) Wild-type AM1 18.5 Quadruple FDH mutant 4.7 Why does the quadruple mutant still consume some HCOO - , since CO 2 production is almost 0? Does this HCOO - go into biomass via the long pathway? HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass
- 33. Biomass flux is not impaired in either the triple or quadruple mutant. My 14 C-Biomass flux data -> HCOO - consumed by the quadruple mutant goes into biomass via the long pathway. (This happens in the wild-type and triple mutant as well.) HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass
- 34. Conclusions on formate dehydrogenase We have now identified all of the major FDHs (4 of them). CH 3 OH can enter the serine cycle via the long pathway, but cannot easily be converted to CO 2 at that point. Thus cells lacking FDH1-4 cannot grow on CH 3 OH alone. HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass FDH 1-4
- 35. Metabolic fluxes in M. extorquens AM1: our view as of 2007 1. The enzyme-mediated long pathway is the cell’s only significant route for assimilating CH 3 OH. 2. Therefore HCOO - , not HCHO, is the key branch point. 3. Formate dehydrogenase (FDH) is important after all! HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass
- 36. Metabolic fluxes in M. extorquens AM1: our view as of 2007 1. The enzyme-mediated long pathway is the cell’s only significant route for assimilating CH 3 OH. 2. Therefore HCOO - , not HCHO, is the key branch point. 3. Formate dehydrogenase (FDH) is important after all! HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass
- 37. Metabolic fluxes in M. extorquens AM1: our view as of 2007 1. The enzyme-mediated long pathway is the cell’s only significant route for assimilating CH 3 OH. 2. Therefore HCOO - , not HCHO, is the key branch point. 3. Formate dehydrogenase (FDH) is important after all! HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass FDH 1-4
- 38. Ongoing and future work C 1 metabolic network • Since HCOO - is the key branch point, study the regulation of FDHs and FtfL. HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass FDH 1-4 FtfL
- 39. Ongoing and future work Interaction of C 1 and multicarbon networks • Two versions of the glyoxylate regeneration cycle have been proposed (Korotkova et al. 2002; Albers et al. 2006). Which occurs in AM1? • How (in terms of enzyme regulation/activity, metabolite levels, and fluxes) do cells transition from succinate (C 4 H 4 O 4 2- ) use to CH 3 OH use? HCOO - CH 2 =H 4 F Serine Cycle CO 2 CH 2 =H 4 MPT H 4 MPT CH 3 OH HCHO H 4 F Biomass
- 40. Undergrads can do this stuff! Accessible techniques • enzyme and metabolite assays • cloning, transformation, PCR • growth assays • mathematical modeling Preliminary success • Dan Yates • Jason Lum
- 41. Summary of metabolism research Long-term goal : Redesign methylotrophs for optimal production of useful chemicals from methanol. A B C k 1 k -1 k 2 k -2 MEASURE fluxes via label tracing MODEL fluxes mathematically REDIRECT fluxes via genetic engineering
- 42. Lidstrom lab members: Mila Chistoserdova, Ph.D. Marina Kalyuzhnaya, Ph.D. Mary Lidstrom, Ph.D. Jonathan Miller, M.S. Betsy Skovran, Ph.D. Tim Strovas, Ph.D. Acknowledgments
- 43. Former lab members: Kelly FitzGerald, Ph.D. (UW Tech Transfer) Xiaofeng Guo, Ph.D. (Brigham & Women’s) Chris Marx, Ph.D. (Harvard) Steve Van Dien, Ph.D. (Genomatica) Julia Vorholt, Ph.D. (ETH Zurich) Additional collaborator: George Kos á ly, Ph.D. (UW Mech Eng) Funding: Kirschstein NRSA (NIH) Acknowledgments
- 44. The End Image: hancockpub.lib.in.us
