Tyler jgi web site fri 1050
•Télécharger en tant que PPT, PDF•
0 j'aime•308 vues
Signaler
Partager
Signaler
Partager
Recommandé
Course: Bioinformatics for Biomedical Research (2014).
Session: 1.3- Genome Browsing, Genomic Data Mining and Genome Data Visualization with Ensembl, Biomart and IGV.
Statistics and Bioinformatisc Unit (UEB) & High Technology Unit (UAT) from Vall d'Hebron Research Institute (www.vhir.org), Barcelona.Genome Browsing, Genomic Data Mining and Genome Data Visualization with Ensem...

Genome Browsing, Genomic Data Mining and Genome Data Visualization with Ensem...VHIR Vall d’Hebron Institut de Recerca
Contenu connexe
Similaire à Tyler jgi web site fri 1050
Course: Bioinformatics for Biomedical Research (2014).
Session: 1.3- Genome Browsing, Genomic Data Mining and Genome Data Visualization with Ensembl, Biomart and IGV.
Statistics and Bioinformatisc Unit (UEB) & High Technology Unit (UAT) from Vall d'Hebron Research Institute (www.vhir.org), Barcelona.Genome Browsing, Genomic Data Mining and Genome Data Visualization with Ensem...

Genome Browsing, Genomic Data Mining and Genome Data Visualization with Ensem...VHIR Vall d’Hebron Institut de Recerca
Similaire à Tyler jgi web site fri 1050 (20)
Practical 7 dna, rna and the flow of genetic information5

Practical 7 dna, rna and the flow of genetic information5
Drug TanzeumDiseasesDiabetesGene and Gene Productglucago.docx

Drug TanzeumDiseasesDiabetesGene and Gene Productglucago.docx
MYSTERY MOLECULE PROJECT, PART II(20 points total)BIO1001.docx

MYSTERY MOLECULE PROJECT, PART II(20 points total)BIO1001.docx
Genome Browsing, Genomic Data Mining and Genome Data Visualization with Ensem...

Genome Browsing, Genomic Data Mining and Genome Data Visualization with Ensem...
Pathema Burkholderia Annotation Jamboree: Prokaryotic Annotation Overview

Pathema Burkholderia Annotation Jamboree: Prokaryotic Annotation Overview
A Genome Sequence Analysis System Built With Hypertable

A Genome Sequence Analysis System Built With Hypertable
Apollo annotation guidelines for i5k projects Diaphorina citri

Apollo annotation guidelines for i5k projects Diaphorina citri
Plus de Sucheta Tripathy
Plus de Sucheta Tripathy (20)
Dernier
💉💊+971581248768>> SAFE AND ORIGINAL ABORTION PILLS FOR SALE IN DUBAI AND ABUDHABI}}+971581248768
+971581248768 Mtp-Kit (500MG) Prices » Dubai [(+971581248768**)] Abortion Pills For Sale In Dubai, UAE, Mifepristone and Misoprostol Tablets Available In Dubai, UAE CONTACT DR.Maya Whatsapp +971581248768 We Have Abortion Pills / Cytotec Tablets /Mifegest Kit Available in Dubai, Sharjah, Abudhabi, Ajman, Alain, Fujairah, Ras Al Khaimah, Umm Al Quwain, UAE, Buy cytotec in Dubai +971581248768''''Abortion Pills near me DUBAI | ABU DHABI|UAE. Price of Misoprostol, Cytotec” +971581248768' Dr.DEEM ''BUY ABORTION PILLS MIFEGEST KIT, MISOPROTONE, CYTOTEC PILLS IN DUBAI, ABU DHABI,UAE'' Contact me now via What's App…… abortion Pills Cytotec also available Oman Qatar Doha Saudi Arabia Bahrain Above all, Cytotec Abortion Pills are Available In Dubai / UAE, you will be very happy to do abortion in Dubai we are providing cytotec 200mg abortion pill in Dubai, UAE. Medication abortion offers an alternative to Surgical Abortion for women in the early weeks of pregnancy. We only offer abortion pills from 1 week-6 Months. We then advise you to use surgery if its beyond 6 months. Our Abu Dhabi, Ajman, Al Ain, Dubai, Fujairah, Ras Al Khaimah (RAK), Sharjah, Umm Al Quwain (UAQ) United Arab Emirates Abortion Clinic provides the safest and most advanced techniques for providing non-surgical, medical and surgical abortion methods for early through late second trimester, including the Abortion By Pill Procedure (RU 486, Mifeprex, Mifepristone, early options French Abortion Pill), Tamoxifen, Methotrexate and Cytotec (Misoprostol). The Abu Dhabi, United Arab Emirates Abortion Clinic performs Same Day Abortion Procedure using medications that are taken on the first day of the office visit and will cause the abortion to occur generally within 4 to 6 hours (as early as 30 minutes) for patients who are 3 to 12 weeks pregnant. When Mifepristone and Misoprostol are used, 50% of patients complete in 4 to 6 hours; 75% to 80% in 12 hours; and 90% in 24 hours. We use a regimen that allows for completion without the need for surgery 99% of the time. All advanced second trimester and late term pregnancies at our Tampa clinic (17 to 24 weeks or greater) can be completed within 24 hours or less 99% of the time without the need surgery. The procedure is completed with minimal to no complications. Our Women's Health Center located in Abu Dhabi, United Arab Emirates, uses the latest medications for medical abortions (RU-486, Mifeprex, Mifegyne, Mifepristone, early options French abortion pill), Methotrexate and Cytotec (Misoprostol). The safety standards of our Abu Dhabi, United Arab Emirates Abortion Doctors remain unparalleled. They consistently maintain the lowest complication rates throughout the nation. Our Physicians and staff are always available to answer questions and care for women in one of the most difficult times in their lives. The decision to have an abortion at the Abortion Cl+971581248768>> SAFE AND ORIGINAL ABORTION PILLS FOR SALE IN DUBAI AND ABUDHA...

+971581248768>> SAFE AND ORIGINAL ABORTION PILLS FOR SALE IN DUBAI AND ABUDHA...?#DUbAI#??##{{(☎️+971_581248768%)**%*]'#abortion pills for sale in dubai@
Dernier (20)
2024: Domino Containers - The Next Step. News from the Domino Container commu...

2024: Domino Containers - The Next Step. News from the Domino Container commu...
Why Teams call analytics are critical to your entire business

Why Teams call analytics are critical to your entire business
Strategies for Landing an Oracle DBA Job as a Fresher

Strategies for Landing an Oracle DBA Job as a Fresher
Apidays Singapore 2024 - Scalable LLM APIs for AI and Generative AI Applicati...

Apidays Singapore 2024 - Scalable LLM APIs for AI and Generative AI Applicati...
AWS Community Day CPH - Three problems of Terraform

AWS Community Day CPH - Three problems of Terraform
Repurposing LNG terminals for Hydrogen Ammonia: Feasibility and Cost Saving

Repurposing LNG terminals for Hydrogen Ammonia: Feasibility and Cost Saving
Apidays Singapore 2024 - Modernizing Securities Finance by Madhu Subbu

Apidays Singapore 2024 - Modernizing Securities Finance by Madhu Subbu
How to Troubleshoot Apps for the Modern Connected Worker

How to Troubleshoot Apps for the Modern Connected Worker
A Beginners Guide to Building a RAG App Using Open Source Milvus

A Beginners Guide to Building a RAG App Using Open Source Milvus
Connector Corner: Accelerate revenue generation using UiPath API-centric busi...

Connector Corner: Accelerate revenue generation using UiPath API-centric busi...
EMPOWERMENT TECHNOLOGY GRADE 11 QUARTER 2 REVIEWER

EMPOWERMENT TECHNOLOGY GRADE 11 QUARTER 2 REVIEWER
Apidays New York 2024 - The value of a flexible API Management solution for O...

Apidays New York 2024 - The value of a flexible API Management solution for O...
+971581248768>> SAFE AND ORIGINAL ABORTION PILLS FOR SALE IN DUBAI AND ABUDHA...

+971581248768>> SAFE AND ORIGINAL ABORTION PILLS FOR SALE IN DUBAI AND ABUDHA...
Automating Google Workspace (GWS) & more with Apps Script

Automating Google Workspace (GWS) & more with Apps Script
Mastering MySQL Database Architecture: Deep Dive into MySQL Shell and MySQL R...

Mastering MySQL Database Architecture: Deep Dive into MySQL Shell and MySQL R...
Tyler jgi web site fri 1050
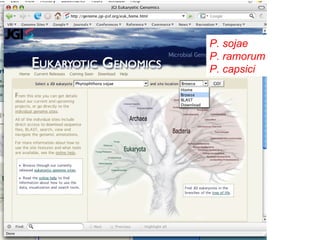
- 1. P. sojae P. ramorum P. capsici
- 2. Detailed manual in use of JGI site
- 4. Genome Browser Can type or paste text in here Then type return or click “Refresh” Change orientation of sequence Display 3 frame translation (must be zoomed in) Display DNA sequence (must be zoomed in) Sequence similarity to P. ramorum Blue = coding Pink = non-coding EVERYTHING IS CLICKABLE!! Click this to access the DNA sequence Tracks give different versions of gene models as well as various sets of Blast hits Clicking on a track will expand it (see next page) Current “best” gene models
- 5. Expanded track Red models have been manually annotated Clicking on open models will open the protein page
- 6. Clicking on model will open the nucleic acid sequence Clicking on green bar will open the protein sequence introns Clicking names will open gene in its database Clicking gray bar will open sequence alignement red blocks are insertions in target Clicking red dot will set sequence as source Protein page EVERYTHING IS CLICKABLE!! Clickable All clickable Blue indicates EST support for UTR
- 7. Shift click or command-click to choose lists of sequences to get (in fasta format) or do Clustal alignments on Shift click or command-click to choose lists of genomes and/or databases for the list of alignments. Then click “Apply Filter” number of alignments to display Protein page OPTIONS
- 8. Sequence Display FROM PROTEIN PAGE Padding controls how much flanking sequence is displayed Introns (should start with GTxxG) Blue indicates EST support for UTR
- 9. Advanced Search (much easier than Simple Search) Shift click or command-click to search mutilple databses simulataneously It’s also an easy way to switch databases Do NOT type return or enter after your entry - you MUST click “Search” Accepts and recognizes almost any kind of query Output of search
- 10. Links: P -> protein page T -> transcript page G -> genome browser Check to download sequences
- 11. Transcript page Contains more info on annotation Used mostly for manual annotation
- 12. Gene Ontology Click to go to protein page
- 13. KEGG Click to go to protein page