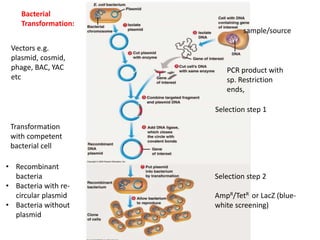
Plant Transformation Techniques and Marker Genes
- 1. Vectors e.g. plasmid, cosmid, phage, BAC, YAC etc sample/source PCR product with sp. Restriction ends, Bacterial Transformation: Selection step 1 Selection step 2 AmpR/TetR or LacZ (blue- white screening) • Recombinant bacteria • Bacteria with re- circular plasmid • Bacteria without plasmid Transformation with competent bacterial cell
- 4. Gene transfer: The process that moves a specific piece of DNA into cell. To introduce genetic diversity into plant population. To select superior plants carrying genes for desired traits. Maintained the range of plant varieties. Genetic transformation: The desirable gene transfer from one organism to another The subsequent stable integration The expression of a foreign gene into the genome. Transgene: The transferred gene Transgenics : The organism that develop after a successful gene transfer Transformed plants: The plants that carry the stably integrated foreign genes. Gene Transfer in Plants
- 5. Transient and stable gene expression Transient expression: The transferred DNA may be expressed for only a short period of time following the DNA transfer process. Transient assay: Analysis of gene expression Rapid monitoring of gene transfer Stable expression DNA is integrated into the plant nuclear or plasmid genome Expression occurs in regenerated plants DNA is inherited in subsequent generation The developmental potential is important, transformed cell should be capable of regenerate into fertile plants.
- 6. Marker genes A set of genes are used for monitoring and detection of plant transformation system in order to know whether the DNA has been successfully transferred into recipient cells. The marker gene is induced along with the target gene. Marker genes are two types- 1. Reporter or scoreable genes (quantitative) 2. Selectable marker genes (qualitative) Easy assay technique, requires no DNA extraction, electrophoresis, or autoradiography.
- 8. Reporter genes Screenable, scoreable, quantitative reporter genes Show immediate expression in cells Analysis of plant gene expression and standardization of parameters for successful gene transfer in a particular technique Should have the following features 1. Detection with high sensitivity 2. Low endogenous background 3. There should be a quantitative assay 4. Assay should be non destructive 5. Assay should be require a minimal amount of effort and expense
- 9. Opine synthase (ocs, nos genes): Reporter genes Enolpyruvyl Shikimate Phosphate Synthase (EPSP): Bar Gene: 2. Herbicide Resistance Genes: 1. Antibiotic Resistance Genes : Neomycin Phosphotransferase II (NPT II): Hygromycin Phosphotransferase (hpt): Selectable Marker Genes Chloramphenicol acetyl transferase (cat gene): Bacterial luciferase (lux F2 genes): Firefly luciferase (luc gene): Green fluorescent protein (gfp gene): β-Glucuronidase (GUS) (uidA gene):
- 10. Opine synthase (ocs, nos genes): octopine and nopaline are common opines in Agrobacterium, respectively produced by the synthase genes ocs and nos. octopine synthase and nopaline synthase genes present in T-DNA of Ti (tumor inducing) or Ri (root inducing) plasmids of Agrobacterium. The transformed status of the plant cells can be easily detected by the presence of these opines. Opines can be separated by electrophoresis and identified. The enzyme activities responsible for the production of opines can also be assayed, enzymes are stable, the enzymatic assay is inexpensive and easy to perform.
- 11. Chloramphenicol acetyl transferase (cat gene): The gene for chloramphenicol acetyl transferase (CAT) is a widely used reporter gene in eukaryotic organism. This gene carries out acetylation of chloramphenicol. The enzymatic assay is very sensitive and relatively simple.
- 12. β-Glucuronidase (GUS) (uidA gene): β-Glucuronidase (GUS) producing from bacterial gene (uidA) is the most commonly used reporter gene in assessing plant transformation. β-Glucuronidase assays are highly sensitive and nonradioactive assay. Quantitative estimation by fluorometric method and qualitative estimation by histochemical assay. enzyme localization can be detected by fluorogenic substrate 4-MUGluc, or chromogenic substrate X-gluc. Quantitative Fluorometric Qualitative histochemical
- 13. Green fluorescent protein (gfp gene): Green fluorescent protein (GFP), coded by gfp gene, is being widely used . Assays of GFP are easier and non-destructive and early screening of even the primary transplants can be done by GFP. Gene for GFP has been isolated from jelly fish Aequorea victoria which is a luminescent organism. GFP emits fluorescence which can be detected under a fluorescent microscope.
- 14. Firefly luciferase (luc gene): The enzyme firefly luciferase, encoded by the gene luc, isolated from north American firefly photinus pyralis. the enzyme catalyses the oxidative decarboxylation of D-luciferin (ATP dependent) which results in the emission of light that can be detected by sensitive luminometers. Bacterial luciferase (lux F2 genes): The bacterial luciferase genes (luxA and luxB) have originated from Vibrio harveyi and V. fischerii.
- 15. Selection of transformed cells is a key factor in successful genetic transformation. Selectable marker genes enable the transformed cells to survive on media containing toxic level of selection agent whereas nontransformed cells get killed. Selectable marker genes such as antibiotic resistance genes, herbicide tolerance genes and several antimetabolite genes are in common practice. The usefulness of a particular resistance marker depends on the characteristics of selection agent the resistance gene the plant material Selectable Marker Genes
- 16. 1. Antibiotic Resistance Genes as Markers: Antibiotic resistance genes used as selectable markers are basically microbial in origin from E.coli . Antibiotics are toxic to plants by inhibiting protein synthesis particularly in the cell organelle like chloroplast. Neomycin Phosphotransferase II (NPT II): NPT II or amino glycoside 3’ –phosphotransferase II enzyme is the most widely used antibiotic resistance gene in plant transformation strategy. The enzyme is encoded by the nptII gene which is derived from Tn5 transposon. It inactivates a number of aminoglycoside antibiotics (protein synthesis inhibitor) such as including kanamycin, neomycin and puromycin. Kanamycin is the most commonly used selective agent.
- 17. Hygromycin Phosphotransferase: The hygromycin phosphotransferase gene (hpt) is derived from E.coli. Hygromycin B is an aminocyclitol antibiotic that kill non transformed cells by blocking protein synthesis during selection process. It is more toxic than kanamycin.
- 18. 2. Herbicide Resistance Marker Genes: Bar Gene: Bar gene encodes phosphinothricin acetyl transferase (PAT), which inactivates herbicides like biolophos, phosphinothricin. This gene has been cloned from two different strains of Streptomyces hygroscopicus and S. viridochromogenes. This is based on damaging of non-transformed tissues by inactivating glutamine synthase (GS) which is required for the assimilation of NH3 and in the regulation of nitrogen metabolism in plants. Inhibition of glutamine synthase results in the accumulation of NH3 , highly toxic to the cell which causes death of plant cells. Bar gene encodes PAT which converts herbicides into a nonherbicidal acetylated form and confers resistance to the cell.
- 19. Enolpyruvyl Shikimate Phosphate Synthase (EPSP): In plants, the enzyme 5-enolpyruvyl-shikimate -phosphate synthase is essensial for biosynthesis of aromatic aminoacids like tyrosine, tryptophan, etc. Glyphosate (N-phosphonomethyl- glycine) is a potential herbicide, is widely used to kill weed plants by competitive inhibitor of enzyme EPSP synthase, and thereby blocking aromatic aminoacid production. In transformation selection technique, mutant EPSP synthase gene is used as herbicide resistant marker. Expression of mutant EPSP synthase gene in transformed tissues is able to survive against glyphosate herbicide included in the culture medium. In contrast, non-transformed cells get killed by inhibition of EPSP synthase enzyme by herbicide.
- 20. Cloning vector
- 21. pCAMBIA1305.1 11846 bp Lac Z alpha kanamycin (R) hygromycin (R) GUSPlus Catalase intron T-BORDER (R) pVS1 Sta T BORDER (L) pBR322 bom NOS polyA CaMV35S polyA 2x CaMV35S promoter CaMV35S promoter pBR322 ori pVS1 rep BamH I Bgl II Bst EII Bst XI EcoR I Hind III Kpn I Nco I Pml I Pst I Sac I Sal I Spe I Xba I Nhe I (2026) Nhe I (5467) Xho I (8910) Xho I (10004) Sma I (11052) Sph I (11083) Binary vector
- 26. Transfer of T-DNA to plant cells.
- 27. The induction of vir genes and T-DNA transfer.