JBEI Research Highlights - March 2019
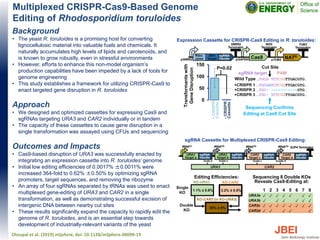
- 1. Multiplexed CRISPR-Cas9-Based Genome Editing of Rhodosporidium toruloides Background • The yeast R. toruloides is a promising host for converting lignocellulosic material into valuable fuels and chemicals. It naturally accumulates high levels of lipids and carotenoids, and is known to grow robustly, even in stressful environments • However, efforts to enhance this non-model organism’s production capabilities have been impeded by a lack of tools for genome engineering • This study establishes a framework for utilizing CRISPR-Cas9 to enact targeted gene disruption in R. toruloides Approach • We designed and optimized cassettes for expressing Cas9 and sgRNAs targeting URA3 and CAR2 individually or in tandem • The capacity of these cassettes to cause gene disruption in a single transformation was assayed using CFUs and sequencing Outcomes and Impacts • Cas9-based disruption of URA3 was successfully enacted by integrating an expression cassette into R. toruloides’ genome • Initial low editing efficiencies of 0.0017% ±0.0011% were increased 364-fold to 0.62% ±0.50% by optimizing sgRNA promoters, target sequences, and removing the ribozyme • An array of four sgRNAs separated by tRNAs was used to enact multiplexed gene-editing of URA3 and CAR2 in a single transformation, as well as demonstrating successful excision of intergenic DNA between nearby cut sites • These results significantly expand the capacity to rapidly edit the genome of R. toruloides, and is an essential step towards development of industrially-relevant variants of the yeast Otoupal et al. (2019) mSphere, doi: 10.1128/mSphere.00099-19 0 50 100 150 Transformantswith GeneDisruption …TGCG TCTCGGTTGACGTG… …TGCGGTCTCGGTTGACGTG… …TGC- -----------GTG… …TGC- TCTCGGTTGACGTG… Wild Type Sequencing Confirms Editing at Cas9 Cut Site P=0.02 -CRISPR Cassette +CRISPR Cassette +CRISPR 1 +CRISPR 2 +CRISPR 3 sgRNA target PAM Cut Site Expression Cassette for CRISPR-Cas9 Editing in R. toruloides: KO-CAR2 (in KO-URA3) 1.1% ± 0.8% 3.2% ± 0.5% KO-URA3 KO-CAR2 30% ± 8% Editing Efficiencies: Single KO Double KO sgRNA Cassette for Multiplexed CRISPR-Cas9 Editing: Sequencing 8 Double KOs Reveals Cas9-Editing at: 1 2 3 4 5 6 7 8 URA3a ✓ ✓ ✓ ✓ ✓ ✓ ✓ ✓ URA3b ✓ ✓ ✓ ✓ ✓ ✓ ✓ ✓ CAR2c ✓ ✓ ✓ ✓ ✓ ✓ ✓ ✓ CAR2d ✓ ✓ ✓ ✓ ✓
- 2. Glycome and proteome components of Golgi membranes are common between two angiosperms with distinct cell wall structures Background • The plant ER-Golgi apparatus is the site of synthesis and assembly of all non-cellulosic polysaccharides, proteoglycans, and proteins destined for the cell wall • As grass species make cell walls distinct from other angiosperms, it has been assumed that the differences in cell wall composition stem from differences in biosynthetic capacities of their respective Golgi Approach • Golgi membranes were prepared from coleoptiles of maize and leaves of Arabidopsis • Glycans in the Golgi preparations were analyzed by immunosorbent-based screens and carbohydrate linkage analysis • Golgi membranes were further purified by free-flow electrophoresis and analyzed by proteomics Outcomes and Impacts • Arabinogalactan-proteins and arabinans represent substantial portions of the Golgi-resident polysaccharides not typically found in high abundance in cell walls of either species • Proteomics yielded over 200 proteins known to function in the biosynthesis and metabolism of cell wall polysaccharides common to all angiosperms, and not just those specific to wall type • We propose that the distinctive compositions of grass primary cell walls result from differential gating or metabolism of secreted polysaccharides post-Golgi by an as yet unknown mechanism, and not by differential expression of genes encoding specific synthase complexes Okekeogbu et al. (2019) Plant Cell, doi: 10.1105/tpc.18.00755. Glycans were analyzed by glycome profiling using specific antibodies. The profiles show many unexpected differences between the Golgi vesicles and the cell walls. For example, xyloglucan is very abundant in maize Golgi but has very low abundance in the wall. The opposite is true for Arabidopsis. Likewise, pectic RGI is very abundant in maize Golgi but has low abundance in the wall.
- 3. Techno‐economic and greenhouse gas analyses of lignin valorization to eugenol and phenolic products in integrated ethanol biorefineries Background • Valorizing lignin is key to improving the economics of lignocellulosic biofuel production • One option for upgrading lignin is to employ catalytic hydrogenolysis assisted by isopropanol (IPA) to convert the lignin recovered after ionic liquid (IL) pretreatment, saccharification, and fermentation • However, a detailed analysis is required to better understand whether this process offers an economic and environmental advantage relative to using lignin for onsite heat and power Approach • Develop a process model for an IL-based biorefinery with lignin upgrading to eugenol, linked with a feedstock availability and logistics model for Mexico • We evaluated the feasibility of this lignin-valorizing biorefinery using locally-available combinations of corn stover, sorghum stubble, and Jatropha fruit shells Outcomes and Impacts • The minimum ethanol selling price (MESP) for this case was $2.02/gal. The resulting cradle‐to‐gate GHG footprint of bioethanol was 21 g CO2‐eq/MJ, a 78% reduction with respect to gasoline when system expansion is used for allocation • Research should target a reduction in IL input by 30% and IPA input by 40%, together with more energy‐efficient separation processes • Sensitivity analysis showed that, for biomass prices higher than $45/t, biorefinery capacities must exceed 5000 t/d biomass input Martinez-Hernandez et al. (2019) Biofuel. Bioprod. Biorefin., doi: 10.1002/bbb.1989)
- 4. The multi-scale challenges of biomass fast pyrolysis and bio-oil upgrading Background • Biomass fast pyrolysis is potentially one of the cheapest routes toward renewable liquid fuels • Crude pyrolysis oil is not immediately usable in the current energy infrastructure, due to undesirable properties such as low energy content and corrosiveness as a result of its high oxygenate content Approach • While various types of pyrolysis reactors and upgrading technologies are under development, knowledge transfer and closing the gap between theory and application requires model development • The conceptual process design including the choice of feedstock, pretreatment technologies, design and operation of pyrolysis and upgrading reactors, and the role of phenomenological modeling in technology transfer was elaborated in this review Outcomes and Impacts • Underpinning economic and environmental impacts of biofuel production requires expanding the system boundaries to include the overall process and supply chain • Design of multi-functional catalysts which can telescope the reaction networks toward high-value fuels and chemicals will enhance the energy conversion efficiency Sharifzadeh et al. (2019) Progress in Energy and Combustion Science, https://doi.org/10.1016/j.pecs.2018.10.006 The multi-scale nature of research into biomass pyrolysis
- 5. Linking ‘omics’ to function unlocks the biotech potential of non-model fungi Background • Non-model fungi are increasingly used in biotechnology, spanning medical, industrial, and agricultural applications, but few have been sequenced compared to yeasts and prokaryotes • This study focuses on the barriers that must be overcome to to translate in silico discoveries enabled by new fungal genomes to applications in biotechnology and biomass hydrolysis Approach • This review highlights recent examples where bioinformatics was used to identify genes and pathways of interest from fungi that were exploited to produce biotechnologically important secondary metabolites, transporters, and lignocellulose-active enzymes Outcomes and Impacts • Implementation of long-read sequencing technologies has accelerated the the number of high-quality fungal genomes and transcriptomes available for study • It is critical to combine sequencing information and systems biology to guide both genetic engineering and heterologous expression strategies to harness the biotech potential of non-model fungi • Direct experimental validation and exploitation of fungal systems lag behind the rate at which novel enzymes and biosynthetic pathways are identified in silico • Additional genome scale models and genome editing techniques must be developed to enable computationally-driven metabolic engineering of non-model fungi Wilken et al. (2019) Current Opinion in Systems Biology, doi: 10.1016/j.coisb.2019.02.001 Vast amount of sequencing data have been recently collected for non-model fungi. Systems and bioinformatic tools are used to identify enzymes and pathways of biotech relevance. In silico analyses and predictions must be coupled with experimental techniques to validate the gene annotation.
- 6. Harnessing Nature’s Anaerobes for Biotechnology and Bioprocessing Background • Industrial application of microbial biotechnology is typically biased toward model microbes with straightforward culturing requirements, genetic engineering tools, and production scaling. • Anaerobes are microorganisms that thrive in the absence of oxygen, and they have widely different culture requirements and a unique metabolism compared to model microbes. • This study considers the opportunities and challenges associated with implementing anaerobes in scalable bioprocessing strategies. Approach • We review current and future uses for anaerobes in biotechnology and bioprocesing in the post-genomics era, with a focus on lignocellulose degradation. Outcomes and Impacts • Early-branching anaerobic fungi (Neocallimastigomycota) have recently been sequenced to reveal a wealth of carbohydrate active enzymes, carbohydrate transporters, and secondary metabolite biosynthesis pathways. • Enzymatic cocktails of secreted enzymes from anaerobic fungi have demonstrated activity on par with many industrial formulations. • Resolving current genetic intractability, scale-up, and cultivation challenges will unlock the potential of lignocellulolytic anaerobes to accelerate bio-based production. Podolsky et al. (2019) Annual Review of Chemical and Biomolecular Engineering., doi: 10.1146/annurev-chembioeng-060718-030340 Anaerobic gut fungi (AGF) biotechnological applications exploit unique features: (a) robust cellulose-, hemicellulose-, and possibly lignin-degrading enzymes; (b) biohydrolysate transporters for heterologous expression; (c) novel secondary metabolite clusters; and (d) lignocellulose fermentation by consortia that lead to the production of sustainable chemicals and fuels.