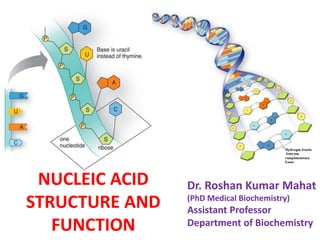
Nucleic acid structure and function
- 1. NUCLEIC ACID STRUCTURE AND FUNCTION Dr. Roshan Kumar Mahat (PhD Medical Biochemistry) Assistant Professor Department of Biochemistry
- 3. Structure of DNA DNA is a polymer of deoxyribonucleoside monophosphates covalently linked by 3'→5'– phosphodiester bonds. With the exception of a few viruses that contain single-stranded (ss) DNA, DNA exists as a double stranded (ds) molecule, in which the two strands wind around each other, forming a double helix.
- 4. In eukaryotic cells, DNA is found associated with various types of proteins (known collectively as nucleoprotein) present in the nucleus, whereas in prokaryotes, the protein– DNA complex is present in a non-membrane bound region known as the nucleoid.
- 5. Purine and pyrimidine bases in DNA
- 6. Figure: Deoxyribose and ribose, the sugars of DNA and RNA. The carbon atoms are numbered from 1 to 5. When the sugar is attached to a base, the carbon atoms are numbered from 1 to 5 to distinguish it from the base. In deoxyribose, the X = H; in ribose, the X =OH.
- 7. 3'→5'-Phosphodiester bonds: Phosphodiester bonds join the 3'-hydroxyl group of the deoxy pentose of one nucleotide to the 5'- hydroxyl group of the deoxy pentose of an adjacent nucleotide through a phosphate group. The resulting long, unbranched chain has polarity, with both a 5'-end and a 3'-end that are not attached to other nucleotides.
- 9. Phosphodiester linkages between nucleotides (in DNA or RNA) can be cleaved hydrolytically by chemicals, or hydrolyzed enzymatically by a family of nucleases: deoxyribonucleases for DNA and ribonucleases for RNA. [Note: Only RNA is cleaved by alkali.]
- 10. Concept of base paring: Figure: Base pairs of DNA
- 11. As a consequence of base pairing, the two strands of DNA are complementary; that is, adenine on one strand corresponds to thymine on the other strand, and guanine corresponds to cytosine. Figure: Two complementary DNA sequences. Hydrogen bonds, plus the hydrophobic interactions between the stacked bases, stabilize the structure of the double helix.
- 12. Note: The specific base pairing in DNA leads to the Chargaff Rule: In any sample of dsDNA, the amount of adenine equals the amount of thymine, the amount of guanine equals the amount of cytosine, and the total amount of purines equals the total amount of pyrimidines.
- 13. The concept of base pairing proved to be essential for determining the mechanism of DNA replication and the mechanisms of transcription and translation. Base pairing allows one strand of DNA to serve as a template for the synthesis of the other strand. Base pairing also allows a strand of DNA to serve as a template for the synthesis of a complementary strand of RNA.
- 14. FIG: DNA strands serve as templates. During replication, the strands of the helix separate in a localized region. Each parental strand serves as a template for the synthesis of a new DNA strand.
- 15. DNA Strands Are Antiparallel: Fig: Antiparallel strands of DNA. This concept of directionality of nucleic acid strands is essential for understanding the mechanisms of replication and transcription.
- 16. The Double Helix: Because each base pair contains a purine bonded to a pyrimidine, the strands are equidistant from each other throughout. If two strands that are equidistant from each other are twisted at the top and the bottom, they form a double helix. In the double helix of DNA, the base pairs that join the two strands are stacked like a spiral staircase along the central axis of the molecule.
- 17. The spatial relationship between the two strands in the helix creates a major (wide) groove and a minor (narrow) groove. These grooves provide access for the binding of regulatory proteins to their specific recognition sequences along the DNA chain. Note: Certain anticancer drugs, such as dactinomycin (actinomycin D), exert their cytotoxic effect by intercalating into the narrow groove of the DNA double helix, thus interfering with DNA and RNA synthesis.
- 18. Fig: Two DNA strands twist to form a double helix.
- 19. Structural forms of double helix The DNA molecule exist in various forms depending on the base composition and physical conditions: 1. The A form 2. the B form, described by Watson and Crick 3. The C form 4. The D form 5. the E form, and 6. the Z form.
- 20. Property A-DNA B-DNA Z-DNA Type of helix Right handed Right handed Left handed Base pairs/turn 11 10 12 Distance between adjacent bases 2.3 A0 3.4 A0 3.8 A0 Helix pitch 28.6 A0 34 A0 44.4 A0 Helical diameter 23 A0 20 A0 18 A0 Major groove Narrow and very deep Wide and deep Wide and flat Minor groove Wide and shallow Narrow and deep Very narrow Comparison of A-DNA, B-DNA and Z-DNA
- 21. Figure: Z, B, and A forms of DNA. The solid red lines connect one phosphate group to the next.
- 22. Denaturation of DNA The term denaturation refers to disruption of native conformation of a biomolecule, so that it does not retain its higher order structure. In case of DNA, denaturation refers to separation of the double strands of DNA into two complementary strands. The double-stranded structure of DNA can be separated into two component strands in solution by increasing the temperature or decreasing the salt concentration.
- 23. Melting temperature: • The strands of a given molecule of DNA separate over a temperature range. • The midpoint is called the melting temperature, or Tm. • When DNA is heated, the temperature at which one half of the helical structure is lost is defined as the melting temperature (Tm).
- 24. The Tm is influenced by the base composition of the DNA and by the salt concentration of the solution. At the physiological pH and ionic strength, the melting temperature of DNA is between 850C to 950C. DNA rich in G–C pairs, which have three hydrogen bonds, melts at a higher temperature than that rich in A–T pairs, which have two hydrogen bonds. For every 10% increase in GC content, the melting temperature increases by 50C.
- 25. Monitoring of strand separation: Denaturation affects properties of DNA and these can be used to monitor strand separation. 1. Hyperchromic effect: the heterocyclic DNA bases absorb UV light of wavelength 260nm. Absorbance of this light increases by 40 % upon denaturation. This is called hyperchromic effect. 2. Viscosity: viscosity of DNA solution decreases on melting because the single strands are far more flexible than the stiff, resilient double helix.
- 26. Figure: Melting temperatures (Tm) of DNA molecules with different nucleotide compositions. (At a wavelength of 260 nm, single- stranded DNA has a higher relative absorbance than does double- stranded DNA.)
- 27. Denatured DNA can undergo renaturation: Under appropriate conditions, complementary DNA strands can reform the double helix by the process called renaturation (or reannealing). It is a rapid process, even the large DNA molecules make take few seconds to minutes for renaturation.
- 28. Figure: Reversible denaturation and annealing (renaturation) of DNA.
- 29. The maximum buoyant density of DNA is 1.70±0.01. The DNA molecule having higher GC content have more compact structure and hence greater buoyant density. For every 10% increase in the GC content, the density rises by 0.12 units. Buoyant density:
- 31. General features of RNA RNA is a polymer of purine and pyrimidine ribonucleotides linked together by 3′,5′- phosphodiester bonds analogous to those in DNA.
- 32. Although sharing many features with DNA, RNA possesses several specific differences: 1. In RNA, the sugar moiety to which the phosphates and purine and pyrimidine bases are attached is ribose rather than the 2′-deoxyribose of DNA. 2. Although RNA contains the ribonucleotides of adenine, guanine, and cytosine, it does not possess thymine except in the rare case. Instead of thymine, RNA contains the ribonucleotide of uracil.
- 33. 3. RNA typically exists as a single strand. However, given the proper complementary base sequence with opposite polarity, the single strand of RNA is capable of folding back on itself like a hairpin and thus acquiring double-stranded characteristics: G pairing with C, and A pairing with U.
- 34. Figure: Diagrammatic representation of the secondary structure of a single-stranded RNA molecule in which a stem loop, or “hairpin,” has been formed.
- 35. 4. Since the RNA molecule is a single strand complementary to only one of the two strands of a gene, its guanine content does not necessarily equal its cytosine content, nor does its adenine content necessarily equal its uracil content. 5. RNA can be hydrolyzed by alkali to 2′, 3′ cyclic diesters of the mononucleotides, compounds that cannot be formed from alkali-treated DNA because of the absence of a 2′-hydroxyl group. The alkali lability of RNA is useful both diagnostically and analytically.
- 36. Types of RNA In all prokaryotic and eukaryotic organisms, three main classes of RNA molecules exist: 1. Messenger RNA(m RNA) 2. Transfer RNA (t RNA) 3. Ribosomal RNA (r RNA) The other are: 1. small nuclear RNA (SnRNA), 2. micro RNA(mi RNA) and 3. small interfering RNA(Si RNA) and 4. heterogeneous nuclear RNA (hnRNA).
- 37. Ribosomal RNA rRNAs are found in association with several proteins as components of the ribosomes— the complex structures that serve as the sites for protein synthesis. There are three distinct size species of rRNA (23S, 16S, and 5S) in prokaryotic cells. In the eukaryotic cytosol, there are four rRNA species (28S, 18S, 5.8S, and 5S). Together, rRNAs make up about 80% of the total RNA in the cell.
- 38. Figure: Comparison of prokaryotic and eukaryotic ribosomes. The cytoplasmic ribosomes of eukaryotes are shown. Mitochondrial ribosomes are similar to prokaryotic ribosomes, but they are smaller (55S rather than 70S).
- 39. Transfer RNA tRNAs are the smallest (4S) of the three major types of RNA molecules. tRNAs serve as adapters for the translation of the information in the sequence of nucleotides of the mRNA into specific amino acids. There are at least 20 species of tRNA molecules in every cell, at least one (and often several) corresponding to each of the 20 amino acids required for protein synthesis.
- 40. The primary structure—that is, the nucleotide sequence—of all tRNA molecules allows extensive folding and intrastrand complementarity to generate a secondary structure that appears in two dimensions like a cloverleaf. tRNAs make up about 15% of the total RNA in the cell. The tRNA molecules contain a high percentage of unusual bases
- 41. tRNA has 5 arms: 1. Acceptor arm- for the attachment of amino acid to form aminoacyl tRNA. 2. D arm 3. TΨC arm 4. Variable arm 5. Anticodon arm: contain sequence of 3 bases that are complementary to codon mRNA. Help define a specific tRNA
- 42. Figure: The tRNA cloverleaf
- 43. Figure: The three-dimensional folding of tRNA.
- 44. Messenger RNA This class is the most heterogeneous in abundance, size and stability. mRNA comprises only about 5% of the RNA in the cell. All members of this RNA class function as messengers conveying the information in a gene to the protein-synthesizing machinery, where each mRNA serves as a template on which a specific sequence of amino acids is polymerized to form a specific protein molecule, the ultimate gene product.
- 45. Figure: The expression of genetic information in DNA into the form of an mRNA transcript with 5’ to 3’ polarity shown.
- 46. Eukaryotic mRNA consists of a leader sequence at the 5-end, a coding region, and a trailer sequence at the 3-end. • The leader sequence begins with a guanosine cap structure at its 5-end. • The coding region begins with a trinucleotide start codon that signals the beginning of translation, followed by the trinucleotide codons for amino acids, and ends at a termination signal.
- 47. The trailer sequence terminates at its 3-end with a poly(A) tail that may be up to 200 nucleotides long. Most of the leader sequence, all of the coding region, and most of the trailer are formed by transcription of the complementary nucleotide sequence in DNA. However, the terminal guanosine in the cap structure and the poly(A) tail do not have complementary sequences; they are added after transcription has been completed (posttranscriptionally).
- 48. Figure: The regions of eukaryotic mRNA. The wavy line indicates the polynucleotide chain of the mRNA and the A’s constituting the poly(A) tail. The 5-cap consists of a guanosine residue linked at its 5-hydroxyl group to three phosphates, which are linked to the 5-hydroxyl group of the next nucleotide in the RNA chain (a 5–5 triphosphate linkage). The start and stop codons represent where protein synthesis is initiated and terminated from this mRNA.
- 49. Figure: The cap structure attached to the 5’ terminal of most eukaryotic messenger RNA molecules.
- 50. The cap is involved in the recognition of mRNA by the translation machinery, and also helps stabilize the mRNA by preventing the nucleolytic attack by 5′-exonucleases. The poly(A) “tail” at the 3′-hydroxyl terminal of mRNAs maintains the intracellular stability of the specific mRNA by preventing the attack of 3′- exonucleases and also facilitate translation.
- 51. Small RNA A large number of discrete, highly conserved, and small RNA species are found in eukaryotic cells; some are quite stable. Most of these molecules are complexed with proteins to form ribonucleoproteins and are distributed in the nucleus, the cytoplasm, or both. They range in size from 20 to 1000 nucleotides and are present in 100,000 to 1,000,000 copies per cell, collectively representing ≤5% of cellular RNA.
- 52. Table: Some of the Species of Small-Stable RNAs Found in Mammalian Cells
- 53. Small Nuclear RNAs (snRNAs) snRNAs, a subset of the small RNAs, are significantly involved in mRNA processing (splicing) and gene regulation. Of the several snRNAs, U1, U2, U4, U5, and U6 are involved in intron removal and the processing of hnRNA into mRNA. The U7 snRNA is involved in production of the correct 3' ends of histone mRNA—which lacks a poly(A) tail.
- 54. Micro RNAs (miRNAs) and Small Interfering RNAs (siRNAs) These two classes of RNAs represent a subset of small RNAs; both play important roles in gene regulation. miRNAs and siRNAs cause inhibition of gene expression by decreasing specific protein production by targeting mRNAs through one of several distinct mechanisms
- 55. Micro RNAs (miRNAs) miRNAs are typically 21–25 nucleotides in length and are generated by nucleolytic processing of the products of distinct genes/transcription units. The small processed mature miRNAs typically hybridize, via the formation of imperfect RNA- RNA duplexes within the 3'- untranslated regions of specific target mRNAs, leading to translation arrest.
- 56. microRNAs, short non-coding RNAs present in all living organisms, have been shown to regulate the expression of at least half of all human genes. These single-stranded RNAs exert their regulatory action by binding messenger RNAs and preventing their translation into proteins.
- 57. Small Interfering RNAs (siRNAs) siRNAs are generated by the specific nucleolytic processing of large dsRNAs that are either produced from other endogenous RNAs, or dsRNAs introduced into the cell, by for example, RNA viruses. These short siRNAs usually form perfect RNA- RNA hybrids with their distinct targets potentially anywhere within the length of the mRNA where the complementary sequence exists.
- 58. Formation of such RNA-RNA duplexes between siRNA and mRNA results in reduced specific protein production because the siRNA-mRNA complexes are degraded by dedicated nucleolytic machinery. some or all of this mRNA degradation occurs in specific organelles termed P bodies.
- 59. Small interfering RNA (siRNA) are 20-25 nucleotide-long double- stranded RNA molecules that have a variety of roles in the cell. siRNA starts off as a double strand RNA, is processed by Dicer and the RISC and then binds to its target mRNA leading to degradation and 'knockdown' of gene expression.
- 60. Significance of miRNAs and siRNAs Both miRNAs and siRNAs represent exciting new potential targets for therapeutic drug development in humans. In addition, siRNAs are frequently used to decrease or "knock-down" specific protein levels in experimental procedures in the laboratory, an extremely useful and powerful alternative to gene-knockout technology.
- 61. Sr. No. RNA DNA 1. Single stranded mainly except when self complementary sequences are there it forms a double stranded structure (Hair pin structure) Double stranded (Except for certain viral DNAs which are single stranded) 2. Ribose is the main sugar The sugar moiety is deoxy ribose 3. Pyrimidine components differ. Thymine is never found(Except tRNA) Thymine is always there but uracil is never found 4. Being single stranded structureIt does not follow Chargaff’s rule It does follow Chargaff's rule. Difference between RNA and DNA
- 62. Sr. No. RNA DNA 5. RNA can be easily destroyed by alkalies to cyclic diesters of mono nucleotides. DNA resists alkali action due to the absence of OH group at 2’ position 6. RNA is a relatively a labile molecule, undergoes easy and spontaneous degradation DNA is a stable molecule. The spontaneous degradation is very 2 slow. The genetic information can be stored for years together without any change. 7. Mainly cytoplasmic, but also present in nucleus (primary transcript and small nuclear RNA) Mainly found in nucleus, extra nuclear DNA is found in mitochondria, and plasmids etc 8. The base content varies from 100- 5000. The size is variable. Millions of base pairs are there depending upon the organism
- 63. Sr. No. RNA DNA 9. There are various types of RNA – mRNA, r RNA, t RNA, Sn RNA, Si RNA, mi RNA and hn RNA. These RNAs perform different and specific functions. DNA is always of one type and performs the function of storage and transfer of genetic information. 10. No variable physiological forms of RNA are found. The different types of RNA do not change their forms There are variable forms of DNA (A to E and Z) 11. RNA is synthesized from DNA, it can not form DNA(except by the action of reverse transcriptase). It can not duplicate (except in certain viruses where it is a genomic material ) DNA can form DNA by replication, it can also form RNA by transcription. 12. Many copies of RNA are present per cell Single copy of DNA is present per cell.