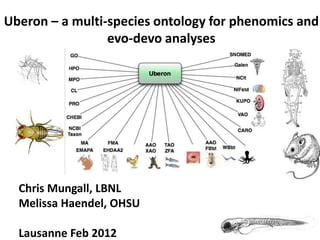
Uberon - A multi-species ontology for phenomics and evo-devo analyses
- 1. Uberon – a multi-species ontology for phenomics and evo-devo analyses Chris Mungall, LBNL Melissa Haendel, OHSU Lausanne Feb 2012
- 2. Outline • Introduction to Bio-Ontologies – Ontologies for data analysis and data integration – Anatomy ontology re-usevsvariation in nature • Uberon – Integration with species anatomy ontologies – Interoperation with non-anatomy ontologies – Reasoning and validation – Handling taxonomic variation – Applications • Homology • Conclusions
- 3. Ontologies abstract over repeated patterns in nature thoracic organ system cavity respiratory thoracic respiratory primordium cavity organ system lung bud lung lung alveolus is_a (SubClassOf) part_of develops_from
- 4. Logical semantics: the difference between ontologies and graphs thoracic organ system cavity respiratory thoracic respiratory primordium cavity organ system lung bud lung lung xinstance-of lung alveolus xinstance-of ‘thoracic cavity organ’ is_a (SubClassOf) part_of develops_from
- 5. Logical semantics: the difference between ontologies and graphs thoracic organ system cavity respiratory thoracic respiratory primordium cavity organ system lung bud lung lung xinstance-of lung alveolus exists y: yinstance-of ‘lung bud’ xdevelops-from y is_a (SubClassOf) part_of develops_from
- 6. Formal semantics allows for more precise queries thoracic organ system cavity respiratory thoracic respiratory primordium cavity organ system lung bud lung xexpressed in y& ypart of z ✔ lung alveolus xexpressed in z Plunc is_a (SubClassOf) part_of xexpressed ubiquitously in y& develops_from ypart of z ✗ expressed in xexpressed ubiquitously in z (inferred)
- 7. Ontology Languages • Web Ontology Language (OWL) – Standard set of logical constructs for building an ontology – Many syntaxes • OWL-RDF/XML • OWL-XML • Manchester – Many reasoners • OBO-Format – Current formalized by mapping to a subset of OWL • can be treated as another OWL syntax
- 8. Many perspectives, many ontologies clinical disorders phenotypes evolutionary characters nervous system anatomy chemical entities cell anatomy gross tissues proteins cells development reactions processes cellular behavior processes physiological processes
- 9. The problem:Data Silos is_a (SubClassOf) part_of GO develops_from surrounded_by FMA multicellularorganism EHDAA2 al process organ system solid organ pharyngeal region respiratory gaseous exchange respiratory primordium respiratory parenchymatous lung bud respiratory system system organ process lung MA Lower thoracic respiratory lobular organ organ system tract cavity MPO abnormal respiratory thoracic respiratory system morphology system cavity organ pleural sac lung abnormal lung morphology lung abnormal pulmonary acinus morphology pulmonary acinus abnormal pulmonary lung alveolus morphology alveolar sac alveolus
- 10. The OBO Foundry • Avoid silo-ization via ontologies that are – open – documented – reusable – interoperable – built according to shared principles – reuse core relations and patterns • Problem: – How do we re-use in the presence of variability? http://obofoundry.org
- 11. Ontologies built for one species will not work for others http://ccm.ucdavis.edu/bcancercd/22/mouse_figure.html http://fme.biostr.washington.edu:8080/FME/index.html
- 12. Generalization leads to complexity cell Variables: V : Variability of entities in domain P : Logical Precision of queries enucleate TP/(TP+FP*c) nucleate cell cell L : “Latticeyness” of class hierarchy ‘exception hierarchy’ Hypothesis: erythrocyte L = kPV nucleate enucleate erythrocyte erythrocyte zebrafish human erythrocyte erythrocyte
- 13. Anatomy Ontology Menagerie • Mouse: Reduced taxonomic scope – MA (adult) – EMAP / EMAPA (embryonic) = • Human Reduced complexity – FMA (adult) – EHDAA2 (CS1-CS20) • Amphibian Historically little – AAO coordination – XAO • Fish – ZFA – TAO • Nematode – WBbt Contrast to: • Arthropod – FBbt (Drosophila) Gene Ontology (GO) – HAO (all kingdoms of life) – Arthropod anatomy ontology
- 14. Sept 2011
- 15. Semantic Similarity of Phenotypes FMA+PATO MP ZFA+PATO FBbt+PATO "Linking Human Diseases to Animal Models Using Ontology-Based Phenotype Annotation." PLoSBiol 7(11): e1000247. doi:10.1371/journal.pbio.1000247 Washington NL, Haendel MA, Mungall CJ, Ashburner M, Westerfield M, Lewis SE
- 16. The problem with mappings Class A Class B In Bioportal? Useful? FMA extensor MA retina Yes No retinaculum of wrist FMA portion of blood MA blood No Yes ZFA Macula MA macula Yes No ZFA aortic arch MA arch of aorta Yes Dubious ZFA hypophysis MA pitiuitary No Yes FMA tibia FBbt tibia Yes No FMA colon GAZColón, Panama Yes No
- 17. Our solution (2008-2009) • Create grouping classes for mappings – Used our own software for entity matching – Manually split/merge in OboEdit using curator knowledge – Internal joke name: Uberon • Used in phenotype analysis – Washington et al – http://owlsim.org • We kept on tweaking – Used for GO logical definitions – Used in cell ontology – Used to clarify and align existing AOs – Integrated logic-based methods – We got criticized, we got better • Fast forward to 2012…
- 18. Uberon in 2012 • Size: – >6500 classes – >19000 relationships (50 relations) – >2000 logical definitions • Scope: – Metazoa • vertebrate bias, in particular mammals • Availability – many versions, in obo and owl • http://uberon.org – Source version is obo, compiled to owl using Oort • What does it look like?
- 19. is_a (SubClassOf) anatomical part_of structure develops_from capable_of is_a (taxon equivalent) endoderm only_in_taxon organ part foregut swim bladder organ endoderm of forgut NCBITaxon: respiration organ Actinopterygii respiratory primordium GO: respiratory gaseous exchange pulmonary acinus alveolus lung lung primordium NCBITaxon: Mammalia alveolus of lung alveolar sac lung bud FMA: pulmonary FMA:lung MA:lung alveolus EHDAA: MA:lung alveolus lung bud Uberon classes generalize species-specific ones, and connect to other ontologies via a variety of relations
- 20. Inter-ontology bridging axioms • Equivalence axioms: – lung (FMA) EquivalentTolung (Ubr) and ‘part of’ some NCBITaxon_9606 – lung (MA) EquivalentTolung (Ubr) and ‘part of’ some NCBITaxon_10090 • Subclass axioms: – lung (EMAPA) SubClassOf lung (Ubr) • Axioms are maintained as xrefs – Translated to full axioms in obo2owl translation (header tags)
- 21. Import closure of Uberon ‘collector’ ontologies
- 22. Different ontology modules ontology contents basic simple relationships uberon main ontology merged main ontology + links to GO, CL, NCBITaxon, NBO taxon collected merged basic metazoan ✔ ✔ vertebrate ✔ ✔ amniote ✔ aves ✔ euarchontoglires ✔
- 23. Collected vs Merged somite somite (Ubr) (Ubr) [includes ZFA axioms as GCIs] somite (ZFA) somite 1 somite 1 (ZFA) (ZFA)
- 24. Logical definitions in GO using Uberon GO:notochord formation: The formation of the notochord from the chordamesoderm. The notochord is composed of large cells packed within a firm connective tissue sheath and is found in all chordates at the ventral surface of the neural tube. In vertebrates, the notochord contributes to the vertebral column. Cross-Product Extensions of the Gene Ontology Journal of Biomedical Informatics 2010. Christopher J. Mungall and Michael Bada and Tanya Z. Berardini and Jennifer Deegan and Amelia Ireland and Midori A. Harris and David P. Hill and Jane Lomax
- 25. Uberon and phenotype ontologies UBERON: retinal blood vessel is_a MA: retina MA:blood vessel FMA:central retinal artery inheres in inheres in MP:abnormal retinal blood vessel HP: Central retinal artery vascular morphology tortuosity
- 26. Logical definitions in CL using Uberon UBERON: epithelium part_of UBERON: trachea CL: epithelial cell part_of is_a CL: tracheal epithelial cell Uberon trachea: A trachea held open by up to 20 C-shaped rings of cartilage. The trachea is the portion of the airway that attaches to the bronchi as it branches. Terrence Meehan, Anna Maria Masci, Amina Abdulla, Lindsay Cowell, Judith Blake, C J Mungall, Alexander Diehl (2011) Logical Development of the Cell Ontology, 6. In BMC Bioinformatics 12 (1)
- 27. Logical definitions in Uberon using external ontologies UBERON: respiratory system capable_of GO: respiratory gaseous exchange part_of UBERON: respiratory airway is_a UBERON: trachea CL: epithelial cell part_of is_a CL: tracheal epithelial cell Uberon logical definitions represent functional, developmental, spatial, etc., axes of classification
- 28. Uberontaxon constraints UBERON: respiratory system capable_of GO: respiratory gaseous exchange part_of UBERON: respiratory airway is_a only_in_taxon Vertebrata UBERON: trachea CL: epithelial cell part_of is_a CL: tracheal epithelial cell J Deegan, E Dimmer, C J Mungall (2010) Formalization of taxon-based constraints to detect inconsistencies in annotation and ontology development, 530. In BMC bioinformatics 11 (1).
- 29. Axioms encoded in OWL provide explicit semantics
- 30. Uberon iterative development cycle Text matching Stem and synonym matching Reasoning Curation • Keep axioms that are manual adding of new classes consistent across AOs obsoletion, merging, splitting • automated consistency checks for disjointeness violations
- 31. Using reasoners to detect errors only_in_taxon UBERON: bone Vertebrata disjoint with is_a is_a Drosophila melanogaster UBERON: tibia Homo sapiens is_a is_a ✗ part_of part_of Fruit fly FBbt‘tibia’ Human FMA ‘tibia’ Developmental Biology, Scott Gilbert, 6th ed.
- 32. Spatial disjointness axioms • Example: – (part_of some midbrain) DisjointWith (part_of some hindbrain) • Note: part_of implies all parts are part of – Brain spatial axioms derived from ABA – Used to find problems in existing mouse ontologies
- 33. Ontology alignment Differences in bone and bone tissue representation
- 34. Ontology alignment Using Uberon for alignment facilitates identification of missing classes
- 35. Managing variation: named subtypes • ‘mammary gland’ part of some ‘female thoracic region’ – humans ✔ – other mammals ✗ • Solution: – mammary gland • thoracic mammary gland • abdominal mammary gland • inguinal mammary gland
- 36. Managing variation: general axioms • adenohypophysis develops from some ‘Rathke’s pouch’ – tetrapoda✔ – teleost✗ • Named subtypes solution – ‘Rathke’s pouch-derived adenohypophysis’ • ugly! • Alternative: – use anonymous classes / OWL general axioms: • (adenohypohysis and part of some tetrapoda) develops from some ‘Rathke’s pouch’ the adenohypophysis has different developmental origins in different species - while in most basal fish and tetrapods the adenohypophyseal anlagen invaginates to form Rathke’s pouch, in teleost fish the adenohypophysealplacode does not invaginate but rather maintains its initial organization forming a solid structure in the head
- 37. Pharyngeal derivatives • Pharyngeal pouches 1-5 – dorsal and ventral parts • Give rise to different structures in different clades – E.g. • parathyroid from ventral pouch 3 & 4 in many vertebrates • in humans, from dorsal pouches 3 and 4 – Kardong, Vertebrates • All encoded in Uberon using general axioms
- 38. A logic for developmental relationships • Most AOs use a single generic develops from relationship – FBbt distinguishes between direct and transitive development – EHDAA2 includes ‘develops in’ • Different structures give different contributions – E.g. neural crest – Modeled explicitly in EHDAA2 • develops from relationships at very specific leaf nodes • Relation composition – has_partodevelops_from - >has_developmental_contribution_from Credit: Osumi-Sutherland, Haendel and Bard
- 39. Provenance for relations [Term] id: UBERON:0005562 name: thymus primordium … relationship: has_developmental_contribution_from UBERON:0010023 {gci_relation="part_of", gci_filler="NCBITaxon:7778", notes="Elasmobranchii", source="ISBN10:0073040584-table13.1"} ! dorsal part of pharyngeal pouch 2 OWL: (‘thymus primordium’ and part_of some NCBITaxon_7778) SubClassOfhas_developmental_contribution_from ‘dorsal part of pharyngeal pouch 2’ Annotations: source "ISBN10:0073040584-table13.1"
- 40. Use of Uberon enhances species- specific ontologies • Many ontologies lack develops from relationships – mouse • MA ✗ • EMAPA ✗ – human • FMA ✗ • EHDAA2 ✔ • SNOMED-CT ✗ • These can be enhanced by the develops from relationships in uberon – E.g find all pharyngeal arch derivatives • Combine with Bgee expression data for powerful queries – E.g compare gene expression patterns for pharyngeal arch derivatives
- 41. Use of Uberon as building block for other ontologies • Basic science – CL – GO – NBO (behavior) – Phenotype (MP, HP) • Applications – OBI – eagle i
- 42. Applications of Uberon in bioinformatics analyses • Crucial lynchpin in a number of phenotype analyses – Washington, Haendel et al – Mousefinder – Phenomenet • Expression analyses – FANTOM5
- 43. Uberon and homology • Uberon classes do not need to be homologous • We try to state necessary and sufficient conditions for all classes – Genus: parent class – Differentia may be any mix of: • Locational • Histological • Structural • Functional • Developmental • Or homology! • This is essentially essentialist – ‘essentialist’ may make evo-devo folks uncomfortable, but it’s how most ontologies work
- 44. Eyes • Eye: organ and has function in go:visual perception – Compound eye: has part ommatidia – Camera-type eye : equivalent to vHOG eye • vertebrate-type* • cephalod-type* *Not yet in ontology
- 45. adrenal gland – interrenal gland • Single class in vHOG • Distinct classes in Uberon – Score highly on semantic similarity measures do to has_part relationships to cell types • Homology can be handled separately • Open question: – interrenal gland vs bodies? – Homology at the level of gland or cortex?
- 46. Using Uberon and vHOG together • UberHOG?
- 47. Using Uberon and vHOG together • vHUG?
- 48. Proposal • Separation of concerns – essentialist definitions – homology relationships • Create ‘homology knowledgebase’ – Statements anchored to Uberon classes • E.g – lung (Ubr) has property: homologous, has_evidence … – head kidney + bone marrow, has property: homologous, has_evidence… – Use homology ontology – Contributions from vHOG and Phenoscape • Automatically aggregate for powerful queries
- 49. Conclusions Anatomy ontologies have been developed independently and do not integrate well without additional help •Uberon generalizes over species-specific anatomy classes • Includes detailed anatomical knowledge via a variety of relationships •designed for reasoning • Highly interconnected with other ontologies • Homology is largely separated • Growing number of applications •For more info: •http://uberon.org http://genomebiology.com/2012/13/1/R5
- 50. Acknowledgments • Uberon • Ontologies • Jonathan Bard (EHDAA2) • Melissa Haendel • Terry Meehan (CL) • George Gkoutos • Alex Diehl (CL) • Carlo Torniai • Terry Hayamizu (MA/CL) • OnardMejino (FMA) • Suzanna Lewis • David Hill (GO) • David Osumi Sutherland (FBbt/CARO) • Paul Schofield (MPATH) • Wasilla Dahdul (TAO/VAO) • Contributions • Paula Mabee (TAO/VAO) • AlanRuttenberg • Erik Segerdell (XAO) • RobHoehndorf • Monte Westerfield (ZFA) • Cynthia Smith (MP) • WacekKusnierczyk • Maryanne Martone (NIF) • Harry Hochheiser • Frederic Bastian (vHOG) • Marc Robinson-Rechavi (vHOG)
Notes de l'éditeur
- A graph in itself has no semantics. Ontology languages such as OWL provide this
- A graph in itself has no semantics. Ontology languages such as OWL provide this
- A graph in itself has no semantics. Ontology languages such as OWL provide this
- UBERON uses GO or other external ontologies for logicaldefinitions (e.g. chemosensory organ, respiration organ, reproductivesystem -- GO; smooth muscle tissue - CL)
- Point here is that Uberon can help ensure correct usage of gross anatomical terms in CL. In this example, this means that the tracheal epithelial cell should be defined only for Vertebrata. [Term]id: CL:0000307 ! tracheal epithelial cellintersection_of: CL:0000066 ! epithelial cellintersection_of: part_of UBERON:0003126 ! trachea
- UBERON uses GO or other external ontologies for logicaldefinitions (e.g. chemosensory organ, respiration organ, reproductivesystem -- GO; smooth muscle tissue - CL)
- UBERON uses GO or other external ontologies for logicaldefinitions (e.g. chemosensory organ, respiration organ, reproductivesystem -- GO; smooth muscle tissue - CL)
- Uberon uses the only_in_taxon method to make relationships such as lactifierous gland only in taxonMammalia and boneonly in taxon Vertebrata. These relations are useful for human users of the ontology, and can be used forconsistency checking within the ontology. For example if the FBbt class “tibia” (representing a segment ofan insect leg) were accidentally placed as a child of UBERON:0000979 tibia, this would be flagged by thereasoner because tibia is a bone, bones are found only in vertebrates, and FBbt is a Drosophila ontology
