Eukaryotic promoter sequence/Elements
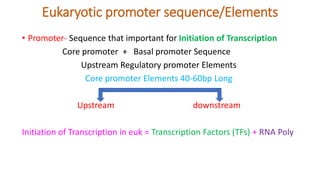
- 1. Eukaryotic promoter sequence/Elements • Promoter- Sequence that important for Initiation of Transcription Core promoter + Basal promoter Sequence Upstream Regulatory promoter Elements Core promoter Elements 40-60bp Long Upstream downstream Initiation of Transcription in euk = Transcription Factors (TFs) + RNA Poly
- 2. RNA Polymerase II Promoter • Upstream of Transcription start site • Core promoter 40-60 bp long 1.BRE= TFIIB Recognizing Elements, 2. MTE= Motif Ten Elements 3. DPE= Downstream promoter Elements, 4.DCE= Downstream core elements • Regulatory Elements = CAAT Box , GC- Box
- 3. RNA Polymerase II mediated Transcription • RNA Polymerase as well as TF Required for Initiation of Transcription. • TF – Trans acting soluble protein Regulation of Transcription • Basal (General)Transcription Factors = TFIID, TFIIA etc Transcription Initiation complex • Regulatory Transcription Factors = Binds to Regulatory sequence Enhancers and silencer Promote or repress the transcription .
- 4. Regulatory Transcription Factors • CBF ( CAAT Binding factors) - Binds to CAAT Box • C/EBP ( CAAT/Enhancer binding protein) Bind to CAAT Box • CREB9 cAMP Response element binding protein) Bind cAMP Response elements
- 6. TFIID – TATA Binding protein (TBP) . Recognized the core promoter. TFIIA- Stabilizes to TBP(TFIID) TFIIB – allow to RNA Pol , core promoter elements TFIIF- Recruitment of RNA Pol II TFIIE- Recruitment of TFII H. TFII H– ATPase and Kinase add –P- On CTD of RNA Pol II to Initiate Transcription.. • Helicase • Regulatory TFs called Activators which bind to specific Regulatory sequence of DNA . • TFs is Coactivator of RNA Pol II mediated Transcription.
- 8. HnRNA • Processing –cleavage • Addition of nucleotides = poly A • Chemical modification – Euk Pro RNA processing nuclear RNA organelle RNA Pre rRNA Pre t RNA
- 9. Transcription Elongation : In Elongation process in eukaryotics includes mRNA processing , 5’ end capping, Splicing, 3’ polyadenylation PTEfB = Protein Transcription Elongation factor B kinase phosphorylation hspTs TATSFI TFIIS 5’ end capping Splicing Hydrolytic Editing and Stimulate to RNA pol. In eukaryotic Transcription in nucleus and translation in cytoplasm so 5’ capping is necessary to prevent degradation of mRNA from cytoplasmic exonuclease enzyme.
- 10. 5’ capping • pTEFb phosphorylates to hspTs which responsible for 5’ capping. • 5’ capping occur simultaneously to the transcription. • 5’ capping is necessary to prevent degradation of 5’ phosphate of mRNA from cytoplasmic exonuclease (lytic)enzyme. • hspTs enzyme have 3 activities RNA tri-phosphatase ,Guanylyl transferase, Guanine N7 Methyl transferase
- 11. RNA tri-phosphatase removes ϒ phosphate from 5’ end of nascent m-RNA. Guanylyl transferase adds α Phosphate on remaining β Phosphate . Guanine N7 Methyl transferase add CH3 group on 7th Nitrogen of GMP at 5’ end . • It is first methylation steps occur in eukaryotic • some higher euk. CH3 group added at second base .
- 12. RNA tri-phosphatase removes ϒ phosphate from 5’ end Guanylyl transferase +GMP Guanine N7 Methyl transferase
- 13. • Now bond formed in between 5’ α Phosphate of GMP and 5’ β Phosphate mRNA is known as 5’5’ bond • Methyl transferase adds CH3( methyl) on N7 of GMP nucleotide. • This is known as 5’ end capping. • Significance of 5’ end capping • It protects mRNA from Degradation • 5’ end capping is necessary for Nuclear export signal (NES) • 5’ end capping is necessary for Translation • It provide stability and transport • Splicing (Pre-mRNA Processing) and 40 s ribosome bind on 5’ cap.
- 14. 3’ Polyadenylation • 250 adenosines add at 3’ end called Polyadenylation • CStF = Cleavage stimulation Factors • CPSF = Cleavage and Polyadenylation Factor • CPSF factor binds on polyadenylation site and CStF bind on GU rich sequence of mRNA. • CF I and CF II breaks mRNA at poly A site . (Cleavage Factor = CF Endonuclease Activity) • After that poly-A- polymerase bind on mRNA poly A site and add AMP on 3’ End. poly-A- polymerase it is template independent RNA Polymerase poly-A- polymerase add 200 nt of adenine on mRNA . • This mRNA is known as Heterogenous mRNA (Hn-RNA) • PABP= Polyadenylation binding protein (poly A tail maintenance protein )
- 15. • Some bacteria having polyadenylation. CF-! And CF-2 PABP= Polyadenylation binding protein (poly A tail maintenance protein ) Poly-A-Polymerase CSTF = Cleavage stimulation Factors CPSF = Cleavage and Polyadenylation Factor
- 16. Significance of Polyadenylation • It protect 3’ end of mRNA From exonuclease • Translation initiation 3’ polyadenylation is necessary.
- 17. Euk. Transcription Termination • Termination of transcription occurs by different process , depending upon the types of RNA Polymerase involves . • RNA Polymerase I / III – stop using the termination factors , Mechanism similar to Rho independent Termination • RNA Polymerase II – How RNA Polymerase II Terminates ? • Two models- allosteric model and Torpedo model 1. Torpedo model –well accepted. • Uncapped 5’ end- it digested by 5’3’ Exonuclease(xnr-2 and Rat-1) * 5’3’ Exonuclease(xnr-2 and Rat-1) digest the 3’ fragment that still associated with the DNA /elongation complex. • Conformation change in RNA Polymerase and get dissociated. 2.Allosteric model – Cleave 3’ Poly-A signal by cleavage factors Conformation change in mRNA And mRNA get dissociate
- 18. Thank You….
